Construct in-house MS2 datbase using metid
Xiaotao Shen (https://www.shenxt.info/)
Created on 2020-03-28 and updated on 2022-09-19
Source:vignettes/database_construction.Rmd
database_construction.RmdIf you have in-house standards which have been acquired with MS2
spectra data, then you can construct the in-house MS2 spectra databases
using the metid package.
There are no specific requirements on how to run the LC/MS data for users. As the in-house database construction in metid is used for users to get the in-house databases for themselves (including m/z, retention time and MS/MS spectra of metabolites, for level 1 annotation (Sumner et al., 2007)), so the users just need to run the standards using the same column, LC-gradient, and MS settings with their real samples in the lab
Data preparation
Firstly, please transform your raw standard MS data (positive and negative modes) to mzXML format using ProteoWizard. The parameter setting is shown in the figure below:
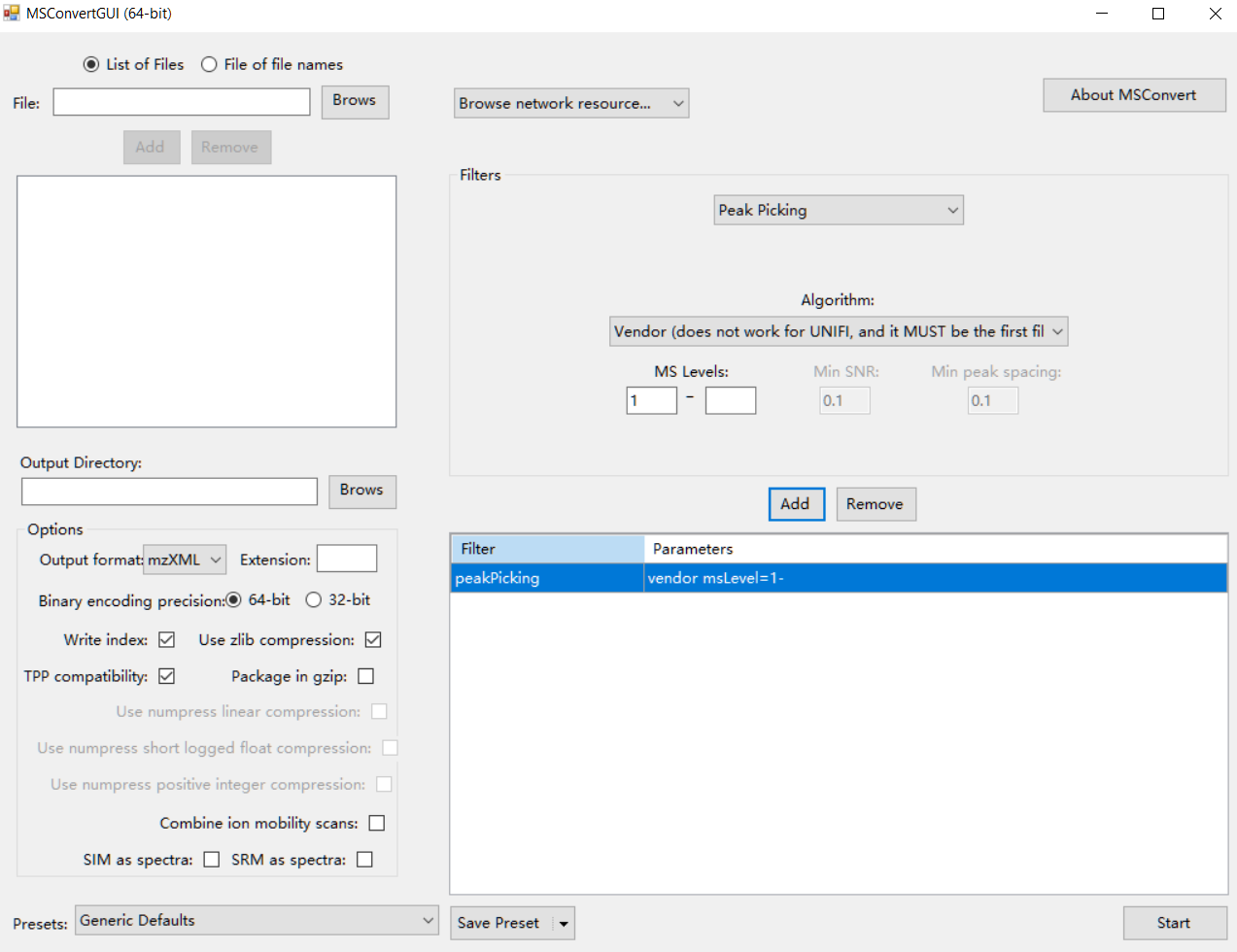
Data organization
Secondly, please organize your standard information as a table, and
output it in a csv or xlsx format. The format of standard information
can refer to our demo data in demoData package.
From column 1 to 11, the columns are “Lab.ID”, “Compound.name”, “mz”, “RT”, “CAS.ID”, “HMDB.ID”, “KEGG.ID”, “Formula”, “mz.pos”, “mz.neg”, “Submitter”, respectively. It is OK if you have other information for the standards. As the demo data show, there are other additional information, namely “Family”, “Sub.pathway” and “Note”.
Lab.ID: No duplicated.
mz: Accurate mass of compounds.
RT: Retention time, unit is second.
mz.pos: Mass to change ratio of compound in positive mode, for example, M+H. You can set it as NA.
mz.neg: Mass to change ratio of compound in negative mode, for example, M-H. You can set it as NA.
Submitter: The name of person or organization. You can set it as NA.
Then create a folder and put your mzXML format datasets (positive
mode in ‘POS’ folder and negative mode in ‘NEG’ folder) and compound
information in it. The mzXML file should have the collision energy in
the name of each file. For example, test_NCE25.mzXML.
The names of the mzXML files should be like this:
xxx_NCE25.mzXML.
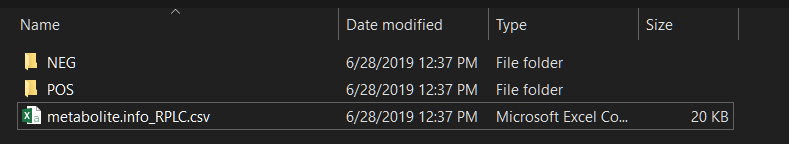
Run construct_database() function
Here we use the demo data from demoData package to show
how to use the construct_database() function to construct
database.
We first prepare dataset.
Download the data here. and then put all of them in the “database_construction” folder.
Then there will be a folder named as
database_construction in your work directory like below
figure shows:
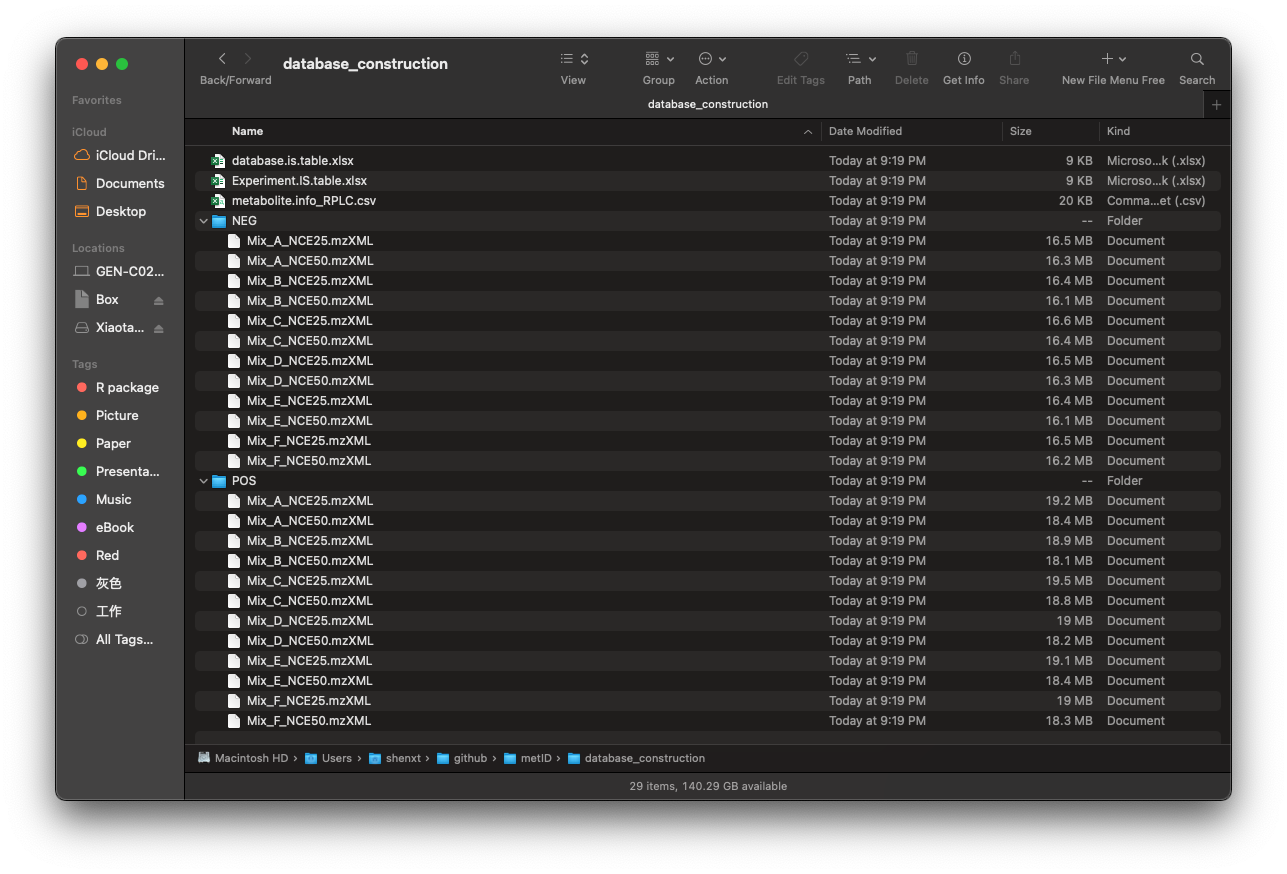 Then we run
Then we run construct_database() function and then we can
get the database.
library(metid)
new.path <- file.path("./database_construction")
test.database <- construct_database(
path = new.path,
version = "0.0.1",
metabolite.info.name = "metabolite.info_RPLC.csv",
source = "Michael Snyder lab",
link = "http://snyderlab.stanford.edu/",
creater = "Xiaotao Shen",
email = "shenxt1990@163.com",
rt = TRUE,
mz.tol = 15,
rt.tol = 30,
threads = 3
)The arguments of construct_database() can be found here
construct_database().
test.database is a databaseClass object, you can print
it to see its information.
test.databaseNote:
test.databaseis only a demo database (metIdentifyClass object). We will don’t use it for next metabolite identification. Then please save this database in you local folder, please note that the saved file name and database name must be same. For example:
save(test.database, file = "test.database")If you save the
test.databaseas a different name, it will be a error when you use it.
MS1 database
If you do not have MS2 data, you can also use
construct_database() function to construct MS1
database.
Session information
sessionInfo()
#> R version 4.2.1 (2022-06-23)
#> Platform: x86_64-apple-darwin17.0 (64-bit)
#> Running under: macOS Big Sur ... 10.16
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] rprojroot_2.0.3 digest_0.6.29 R6_2.5.1 jsonlite_1.8.0
#> [5] magrittr_2.0.3 evaluate_0.15 stringi_1.7.6 rlang_1.0.3
#> [9] cachem_1.0.6 cli_3.3.0 rstudioapi_0.13 fs_1.5.2
#> [13] jquerylib_0.1.4 bslib_0.3.1 ragg_1.2.2 rmarkdown_2.14
#> [17] pkgdown_2.0.5 textshaping_0.3.6 desc_1.4.1 tools_4.2.1
#> [21] stringr_1.4.0 purrr_0.3.4 yaml_2.3.5 xfun_0.31
#> [25] fastmap_1.1.0 compiler_4.2.1 systemfonts_1.0.4 memoise_2.0.1
#> [29] htmltools_0.5.2 knitr_1.39 sass_0.4.1